Geographical Pattern Causality
Wenbo Lyu
Last
update: 2026-04-03
Last run: 2026-04-05
Source: Last run: 2026-04-05
vignettes/main4_gpc.Rmd
main4_gpc.RmdMethodological Background
Geographical Pattern Causality (GPC) infers causal relations from spatial cross-sectional data by reconstructing a symbolic approximation of the underlying spatial dynamical system.
Let \(x(s)\) and \(y(s)\) denote two spatial cross-sections over spatial units \(s \in \mathcal{S}\).
(1) Spatial Embedding
For each spatial unit \(s_i\), GPC constructs an embedding vector
\[ \mathbf{E}_{x(s_i)} = \big( x(s_{i}^{(1)}), x(s_{i}^{(2)}), \dots, x(s_{i}^{(E\tau)}) \big), \]
where \(s_{i}^{(k)}\) denotes the \(k\)-th spatially lagged value of the spatial unit \(s_i\), determined by embedding dimension \(E\) and lag \(\tau\). This yields two reconstructed state spaces \(\mathcal{M}_x, \mathcal{M}_y \subset \mathbb{R}^E\).
(2) Symbolic Pattern Extraction
Local geometric transitions in each manifold are mapped to symbols
\[ \sigma_x(s_i),; \sigma_y(s_i) \in \mathcal{A}, \]
encoding increasing, decreasing, or non-changing modes. These symbolic trajectories summarize local pattern evolution.
(3) Cross-Pattern Mapping
Causality from \(x \to y\) is assessed by predicting:
\[ \hat{\sigma}_y(s_i) = F\big( \sigma_x(s_j): s_j \in \mathcal{N}_k(s_i) \big), \]
where \(\mathcal{N}_k\) denotes the set of \(k\) nearest neighbors in \(\mathcal{M}_x\). The agreement structure between \(\hat{\sigma}_y(s_i)\) and \(\sigma_y(s_i)\) determines the causal mode:
- Positive: \(\hat{\sigma}_y = \sigma_y\)
- Negative: \(\hat{\sigma}_y = -\sigma_y\)
- Dark: neither agreement nor opposition
(4) Causal Strength
The global causal strength is the normalized consistency of symbol matches:
\[ C_{x \to y} = \frac{1}{|\mathcal{S}|} \sum_{s_i \in \mathcal{S}} \mathbb{I}\big[ \hat{\sigma}_y(s_i) \bowtie \sigma_y(s_i) \big], \]
where \(\bowtie\) encodes positive, negative, or dark matching rules.
Usage examples
Example of spatial vector data
Load the spEDM package and its columbus spatial analysis
data:
library(spEDM)
columbus = sf::read_sf(system.file("case/columbus.gpkg", package="spEDM"))
columbus
## Simple feature collection with 49 features and 6 fields
## Geometry type: POLYGON
## Dimension: XY
## Bounding box: xmin: 5.874907 ymin: 10.78863 xmax: 11.28742 ymax: 14.74245
## Projected CRS: Undefined Cartesian SRS with unknown unit
## # A tibble: 49 × 7
## hoval inc crime open plumb discbd geom
## <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <POLYGON>
## 1 80.5 19.5 15.7 2.85 0.217 5.03 ((8.624129 14.23698, 8.5597 14.74245, …
## 2 44.6 21.2 18.8 5.30 0.321 4.27 ((8.25279 14.23694, 8.282758 14.22994,…
## 3 26.4 16.0 30.6 4.53 0.374 3.89 ((8.653305 14.00809, 8.81814 14.00205,…
## 4 33.2 4.48 32.4 0.394 1.19 3.7 ((8.459499 13.82035, 8.473408 13.83227…
## 5 23.2 11.3 50.7 0.406 0.625 2.83 ((8.685274 13.63952, 8.677577 13.72221…
## 6 28.8 16.0 26.1 0.563 0.254 3.78 ((9.401384 13.5504, 9.434411 13.69427,…
## 7 75 8.44 0.178 0 2.40 2.74 ((8.037741 13.60752, 8.062716 13.60452…
## 8 37.1 11.3 38.4 3.48 2.74 2.89 ((8.247527 13.58651, 8.2795 13.5965, 8…
## 9 52.6 17.6 30.5 0.527 0.891 3.17 ((9.333297 13.27242, 9.671007 13.27361…
## 10 96.4 13.6 34.0 1.55 0.558 4.33 ((10.08251 13.03377, 10.0925 13.05275,…
## # ℹ 39 more rowsThe false nearest neighbours (FNN) method helps identify the appropriate minimal embedding dimension for reconstructing the state space of a time series or spatial cross-sectional data.
spEDM::fnn(columbus, "crime", E = 1:10, eps = stats::sd(columbus$crime))
## [spEDM] Output 'E:i' corresponds to the i-th valid embedding dimension.
## [spEDM] Input E values exceeding max embeddable dimension were truncated.
## [spEDM] Please map output indices to original E inputs before interpretation.
## E:1 E:2 E:3 E:4 E:5 E:6 E:7
## 0.59183673 0.04081633 0.04081633 0.10204082 0.00000000 0.00000000 0.00000000
## E:8
## 0.00000000The false nearest neighbours (FNN) ratio decreased to 0
when the embedding dimension E reached 5, and remained relatively stable
thereafter. Therefore, we adopted \(E =
5\) as the minimal embedding dimension for subsequent parameter
search.
Then, search optimal parameters:
# determine the type of causality using correlation
stats::cor.test(columbus$hoval,columbus$crime)
##
## Pearson's product-moment correlation
##
## data: columbus$hoval and columbus$crime
## t = -4.8117, df = 47, p-value = 1.585e-05
## alternative hypothesis: true correlation is not equal to 0
## 95 percent confidence interval:
## -0.7366777 -0.3497978
## sample estimates:
## cor
## -0.5744867
# since the correlation is -0.574, negative causality is selected as the metric to maximize in the optimal parameter search
spEDM::pc(columbus, "crime", "hoval", E = 5:9, k = 7:10, tau = 1, maximize = "negative")
## The suggested E,k,tau for variable hoval is 7, 9 and 1Run geographical pattern causality analysis
spEDM::gpc(columbus, "crime", "hoval", E = 7, k = 9)
## type strength direction
## 1 positive 0.00000000 crime -> hoval
## 2 negative 0.34035656 crime -> hoval
## 3 dark 0.02786836 crime -> hoval
## 4 positive 0.00000000 hoval -> crime
## 5 negative 0.00000000 hoval -> crime
## 6 dark 0.05337361 hoval -> crimeConvergence diagnostics
crime_convergence = spEDM::gpc(columbus, "crime", "hoval",
libsizes = seq(9, 50, by = 1),
E = 7, k = 9, progressbar = FALSE)
crime_convergence
## libsizes type strength direction
## 1 10 positive 0.000000000 crime -> hoval
## 2 11 positive 0.000000000 crime -> hoval
## 3 12 positive 0.000000000 crime -> hoval
## 4 13 positive 0.000000000 crime -> hoval
## 5 14 positive 0.000000000 crime -> hoval
## 6 15 positive 0.000000000 crime -> hoval
## 7 16 positive 0.000000000 crime -> hoval
## 8 17 positive 0.000000000 crime -> hoval
## 9 18 positive 0.000000000 crime -> hoval
## 10 19 positive 0.000000000 crime -> hoval
## 11 20 positive 0.000000000 crime -> hoval
## 12 21 positive 0.000000000 crime -> hoval
## 13 22 positive 0.000000000 crime -> hoval
## 14 23 positive 0.000000000 crime -> hoval
## 15 24 positive 0.000000000 crime -> hoval
## 16 25 positive 0.000000000 crime -> hoval
## 17 26 positive 0.000000000 crime -> hoval
## 18 27 positive 0.000000000 crime -> hoval
## 19 28 positive 0.000000000 crime -> hoval
## 20 29 positive 0.000000000 crime -> hoval
## 21 30 positive 0.000000000 crime -> hoval
## 22 31 positive 0.000000000 crime -> hoval
## 23 32 positive 0.000000000 crime -> hoval
## 24 33 positive 0.000000000 crime -> hoval
## 25 34 positive 0.000000000 crime -> hoval
## 26 35 positive 0.000000000 crime -> hoval
## 27 36 positive 0.000000000 crime -> hoval
## 28 37 positive 0.000000000 crime -> hoval
## 29 38 positive 0.000000000 crime -> hoval
## 30 39 positive 0.000000000 crime -> hoval
## 31 40 positive 0.000000000 crime -> hoval
## 32 41 positive 0.000000000 crime -> hoval
## 33 42 positive 0.000000000 crime -> hoval
## 34 43 positive 0.000000000 crime -> hoval
## 35 44 positive 0.000000000 crime -> hoval
## 36 45 positive 0.000000000 crime -> hoval
## 37 46 positive 0.000000000 crime -> hoval
## 38 47 positive 0.000000000 crime -> hoval
## 39 48 positive 0.000000000 crime -> hoval
## 40 49 positive 0.000000000 crime -> hoval
## 41 10 negative 0.000000000 crime -> hoval
## 42 11 negative 0.000000000 crime -> hoval
## 43 12 negative 0.000000000 crime -> hoval
## 44 13 negative 0.000000000 crime -> hoval
## 45 14 negative 0.000000000 crime -> hoval
## 46 15 negative 0.000000000 crime -> hoval
## 47 16 negative 0.000000000 crime -> hoval
## 48 17 negative 0.000000000 crime -> hoval
## 49 18 negative 0.000000000 crime -> hoval
## 50 19 negative 0.079758388 crime -> hoval
## 51 20 negative 0.095663420 crime -> hoval
## 52 21 negative 0.104704018 crime -> hoval
## 53 22 negative 0.083390936 crime -> hoval
## 54 23 negative 0.155933157 crime -> hoval
## 55 24 negative 0.160468454 crime -> hoval
## 56 25 negative 0.159841305 crime -> hoval
## 57 26 negative 0.153965990 crime -> hoval
## 58 27 negative 0.211917428 crime -> hoval
## 59 28 negative 0.186256958 crime -> hoval
## 60 29 negative 0.174366043 crime -> hoval
## 61 30 negative 0.172561491 crime -> hoval
## 62 31 negative 0.189445238 crime -> hoval
## 63 32 negative 0.212568790 crime -> hoval
## 64 33 negative 0.192410926 crime -> hoval
## 65 34 negative 0.193216971 crime -> hoval
## 66 35 negative 0.192956642 crime -> hoval
## 67 36 negative 0.209433064 crime -> hoval
## 68 37 negative 0.203325807 crime -> hoval
## 69 38 negative 0.203325807 crime -> hoval
## 70 39 negative 0.205654746 crime -> hoval
## 71 40 negative 0.205127658 crime -> hoval
## 72 41 negative 0.209042309 crime -> hoval
## 73 42 negative 0.209042309 crime -> hoval
## 74 43 negative 0.226222782 crime -> hoval
## 75 44 negative 0.226700257 crime -> hoval
## 76 45 negative 0.226904376 crime -> hoval
## 77 46 negative 0.238064699 crime -> hoval
## 78 47 negative 0.312430855 crime -> hoval
## 79 48 negative 0.340356565 crime -> hoval
## 80 49 negative 0.340356565 crime -> hoval
## 81 10 dark 0.000000000 crime -> hoval
## 82 11 dark 0.000000000 crime -> hoval
## 83 12 dark 0.000000000 crime -> hoval
## 84 13 dark 0.000000000 crime -> hoval
## 85 14 dark 0.000000000 crime -> hoval
## 86 15 dark 0.000000000 crime -> hoval
## 87 16 dark 0.000000000 crime -> hoval
## 88 17 dark 0.000000000 crime -> hoval
## 89 18 dark 0.000000000 crime -> hoval
## 90 19 dark 0.000000000 crime -> hoval
## 91 20 dark 0.000000000 crime -> hoval
## 92 21 dark 0.000000000 crime -> hoval
## 93 22 dark 0.000000000 crime -> hoval
## 94 23 dark 0.000000000 crime -> hoval
## 95 24 dark 0.000000000 crime -> hoval
## 96 25 dark 0.000000000 crime -> hoval
## 97 26 dark 0.000000000 crime -> hoval
## 98 27 dark 0.000000000 crime -> hoval
## 99 28 dark 0.000000000 crime -> hoval
## 100 29 dark 0.000000000 crime -> hoval
## 101 30 dark 0.000000000 crime -> hoval
## 102 31 dark 0.000000000 crime -> hoval
## 103 32 dark 0.000000000 crime -> hoval
## 104 33 dark 0.000000000 crime -> hoval
## 105 34 dark 0.000000000 crime -> hoval
## 106 35 dark 0.000000000 crime -> hoval
## 107 36 dark 0.000000000 crime -> hoval
## 108 37 dark 0.000000000 crime -> hoval
## 109 38 dark 0.000000000 crime -> hoval
## 110 39 dark 0.000000000 crime -> hoval
## 111 40 dark 0.000000000 crime -> hoval
## 112 41 dark 0.003273597 crime -> hoval
## 113 42 dark 0.009965908 crime -> hoval
## 114 43 dark 0.017923335 crime -> hoval
## 115 44 dark 0.019163588 crime -> hoval
## 116 45 dark 0.019163588 crime -> hoval
## 117 46 dark 0.026656692 crime -> hoval
## 118 47 dark 0.026656692 crime -> hoval
## 119 48 dark 0.027868360 crime -> hoval
## 120 49 dark 0.027868360 crime -> hoval
## 121 10 positive 0.000000000 hoval -> crime
## 122 11 positive 0.000000000 hoval -> crime
## 123 12 positive 0.000000000 hoval -> crime
## 124 13 positive 0.000000000 hoval -> crime
## 125 14 positive 0.000000000 hoval -> crime
## 126 15 positive 0.000000000 hoval -> crime
## 127 16 positive 0.000000000 hoval -> crime
## 128 17 positive 0.000000000 hoval -> crime
## 129 18 positive 0.000000000 hoval -> crime
## 130 19 positive 0.000000000 hoval -> crime
## 131 20 positive 0.000000000 hoval -> crime
## 132 21 positive 0.000000000 hoval -> crime
## 133 22 positive 0.000000000 hoval -> crime
## 134 23 positive 0.000000000 hoval -> crime
## 135 24 positive 0.000000000 hoval -> crime
## 136 25 positive 0.000000000 hoval -> crime
## 137 26 positive 0.000000000 hoval -> crime
## 138 27 positive 0.000000000 hoval -> crime
## 139 28 positive 0.000000000 hoval -> crime
## 140 29 positive 0.000000000 hoval -> crime
## 141 30 positive 0.000000000 hoval -> crime
## 142 31 positive 0.000000000 hoval -> crime
## 143 32 positive 0.000000000 hoval -> crime
## 144 33 positive 0.000000000 hoval -> crime
## 145 34 positive 0.000000000 hoval -> crime
## 146 35 positive 0.000000000 hoval -> crime
## 147 36 positive 0.000000000 hoval -> crime
## 148 37 positive 0.000000000 hoval -> crime
## 149 38 positive 0.000000000 hoval -> crime
## 150 39 positive 0.000000000 hoval -> crime
## 151 40 positive 0.000000000 hoval -> crime
## 152 41 positive 0.000000000 hoval -> crime
## 153 42 positive 0.000000000 hoval -> crime
## 154 43 positive 0.000000000 hoval -> crime
## 155 44 positive 0.000000000 hoval -> crime
## 156 45 positive 0.000000000 hoval -> crime
## 157 46 positive 0.000000000 hoval -> crime
## 158 47 positive 0.000000000 hoval -> crime
## 159 48 positive 0.000000000 hoval -> crime
## 160 49 positive 0.000000000 hoval -> crime
## 161 10 negative 0.000000000 hoval -> crime
## 162 11 negative 0.000000000 hoval -> crime
## 163 12 negative 0.000000000 hoval -> crime
## 164 13 negative 0.000000000 hoval -> crime
## 165 14 negative 0.000000000 hoval -> crime
## 166 15 negative 0.000000000 hoval -> crime
## 167 16 negative 0.000000000 hoval -> crime
## 168 17 negative 0.000000000 hoval -> crime
## 169 18 negative 0.000000000 hoval -> crime
## 170 19 negative 0.000000000 hoval -> crime
## 171 20 negative 0.000000000 hoval -> crime
## 172 21 negative 0.000000000 hoval -> crime
## 173 22 negative 0.000000000 hoval -> crime
## 174 23 negative 0.000000000 hoval -> crime
## 175 24 negative 0.000000000 hoval -> crime
## 176 25 negative 0.000000000 hoval -> crime
## 177 26 negative 0.000000000 hoval -> crime
## 178 27 negative 0.000000000 hoval -> crime
## 179 28 negative 0.000000000 hoval -> crime
## 180 29 negative 0.000000000 hoval -> crime
## 181 30 negative 0.000000000 hoval -> crime
## 182 31 negative 0.000000000 hoval -> crime
## 183 32 negative 0.000000000 hoval -> crime
## 184 33 negative 0.000000000 hoval -> crime
## 185 34 negative 0.000000000 hoval -> crime
## 186 35 negative 0.000000000 hoval -> crime
## 187 36 negative 0.000000000 hoval -> crime
## 188 37 negative 0.000000000 hoval -> crime
## 189 38 negative 0.000000000 hoval -> crime
## 190 39 negative 0.000000000 hoval -> crime
## 191 40 negative 0.000000000 hoval -> crime
## 192 41 negative 0.000000000 hoval -> crime
## 193 42 negative 0.000000000 hoval -> crime
## 194 43 negative 0.000000000 hoval -> crime
## 195 44 negative 0.000000000 hoval -> crime
## 196 45 negative 0.000000000 hoval -> crime
## 197 46 negative 0.000000000 hoval -> crime
## 198 47 negative 0.000000000 hoval -> crime
## 199 48 negative 0.000000000 hoval -> crime
## 200 49 negative 0.000000000 hoval -> crime
## 201 10 dark 0.050816175 hoval -> crime
## 202 11 dark 0.043383185 hoval -> crime
## 203 12 dark 0.041382131 hoval -> crime
## 204 13 dark 0.040519557 hoval -> crime
## 205 14 dark 0.048272151 hoval -> crime
## 206 15 dark 0.040230136 hoval -> crime
## 207 16 dark 0.044464149 hoval -> crime
## 208 17 dark 0.044665447 hoval -> crime
## 209 18 dark 0.042914041 hoval -> crime
## 210 19 dark 0.041468145 hoval -> crime
## 211 20 dark 0.038849769 hoval -> crime
## 212 21 dark 0.038819148 hoval -> crime
## 213 22 dark 0.038858502 hoval -> crime
## 214 23 dark 0.039451135 hoval -> crime
## 215 24 dark 0.040338936 hoval -> crime
## 216 25 dark 0.038705221 hoval -> crime
## 217 26 dark 0.036890764 hoval -> crime
## 218 27 dark 0.037987604 hoval -> crime
## 219 28 dark 0.039793512 hoval -> crime
## 220 29 dark 0.037765229 hoval -> crime
## 221 30 dark 0.038233561 hoval -> crime
## 222 31 dark 0.036447303 hoval -> crime
## 223 32 dark 0.039415083 hoval -> crime
## 224 33 dark 0.037622935 hoval -> crime
## 225 34 dark 0.038474108 hoval -> crime
## 226 35 dark 0.037545521 hoval -> crime
## 227 36 dark 0.041351817 hoval -> crime
## 228 37 dark 0.039936088 hoval -> crime
## 229 38 dark 0.044526449 hoval -> crime
## 230 39 dark 0.041276856 hoval -> crime
## 231 40 dark 0.044994931 hoval -> crime
## 232 41 dark 0.044496909 hoval -> crime
## 233 42 dark 0.049654135 hoval -> crime
## 234 43 dark 0.050765691 hoval -> crime
## 235 44 dark 0.052651619 hoval -> crime
## 236 45 dark 0.053726028 hoval -> crime
## 237 46 dark 0.053726628 hoval -> crime
## 238 47 dark 0.053373606 hoval -> crime
## 239 48 dark 0.053373606 hoval -> crime
## 240 49 dark 0.053373606 hoval -> crime
plot(crime_convergence, ylimits = c(-0.01,0.61),
xlimits = c(9,50), xbreaks = seq(10, 50, 5))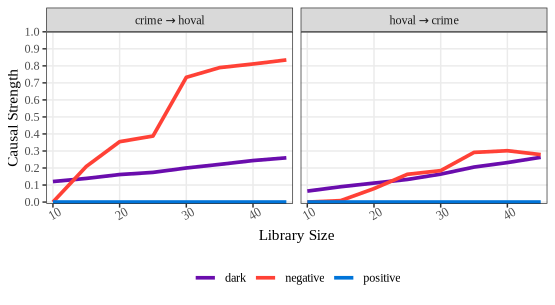
Example of spatial raster data
Load the spEDM package and its farmland NPP data:
library(spEDM)
npp = terra::rast(system.file("case/npp.tif", package = "spEDM"))
# To save the computation time, we will aggregate the data by 3 times
npp = terra::aggregate(npp, fact = 3, na.rm = TRUE)
npp
## class : SpatRaster
## size : 135, 161, 5 (nrow, ncol, nlyr)
## resolution : 30000, 30000 (x, y)
## extent : -2625763, 2204237, 1867078, 5917078 (xmin, xmax, ymin, ymax)
## coord. ref. : CGCS2000_Albers
## source(s) : memory
## names : npp, pre, tem, elev, hfp
## min values : 187.50, 390.3351, -47.8194, -110.1494, 0.04434316
## max values : 15381.89, 23734.5330, 262.8576, 5217.6431, 42.68803711
# Check the validated cell number
nnamat = terra::as.matrix(npp[[1]], wide = TRUE)
nnaindice = which(!is.na(nnamat), arr.ind = TRUE)
dim(nnaindice)
## [1] 6920 2Determining minimal embedding dimension:
spEDM::fnn(npp, "npp", E = 1:15,
eps = stats::sd(terra::values(npp[["npp"]]),na.rm = TRUE))
## [spEDM] Output 'E:i' corresponds to the i-th valid embedding dimension.
## [spEDM] Input E values exceeding max embeddable dimension were truncated.
## [spEDM] Please map output indices to original E inputs before interpretation.
## E:1 E:2 E:3 E:4 E:5 E:6
## 0.9813070569 0.5309427415 0.1322254335 0.0167630058 0.0017341040 0.0000000000
## E:7 E:8 E:9 E:10 E:11 E:12
## 0.0001445087 0.0000000000 0.0000000000 0.0000000000 0.0002890173 0.0000000000
## E:13 E:14
## 0.0001445087 0.0001445087At \(E = 6\), the false nearest neighbor ratio stabilizes approximately at 0.0001 and remains constant thereafter. Therefore, \(E = 6\) is selected as minimal embedding dimension for the subsequent GPC analysis.
Then, search optimal parameters:
stats::cor.test(~ pre + npp,
data = terra::values(npp[[c("pre","npp")]],
dataframe = TRUE, na.rm = TRUE))
##
## Pearson's product-moment correlation
##
## data: pre and npp
## t = 108.74, df = 6912, p-value < 2.2e-16
## alternative hypothesis: true correlation is not equal to 0
## 95 percent confidence interval:
## 0.7855441 0.8029426
## sample estimates:
## cor
## 0.7944062
g1 = spEDM::pc(npp, "pre", "npp", E = 6:10, k = 7:12, tau = 1:5, maximize = "positive")
g1
## The suggested E,k,tau for variable npp is 6, 9 and 5Run geographical pattern causality analysis
spEDM::gpc(npp, "pre", "npp", E = 6, k = 9, tau = 5)
## type strength direction
## 1 positive 0.5222514 pre -> npp
## 2 negative 0.3180282 pre -> npp
## 3 dark 0.3795653 pre -> npp
## 4 positive 0.4491962 npp -> pre
## 5 negative 0.6663956 npp -> pre
## 6 dark 0.4303680 npp -> preConvergence diagnostics
npp_convergence = spEDM::gpc(npp, "pre", "npp",
libsizes = matrix(rep(seq(10,80,10),2), ncol = 2),
E = 6, k = 9, tau = 5, progressbar = FALSE)
npp_convergence
## libsizes type strength direction
## 1 100 positive 0.13573984 pre -> npp
## 2 400 positive 0.18147283 pre -> npp
## 3 900 positive 0.24614666 pre -> npp
## 4 1600 positive 0.33710096 pre -> npp
## 5 2500 positive 0.40395843 pre -> npp
## 6 3600 positive 0.45524137 pre -> npp
## 7 4900 positive 0.48874459 pre -> npp
## 8 6400 positive 0.51020746 pre -> npp
## 9 100 negative 0.00000000 pre -> npp
## 10 400 negative 0.00000000 pre -> npp
## 11 900 negative 0.00000000 pre -> npp
## 12 1600 negative 0.00000000 pre -> npp
## 13 2500 negative 0.05496496 pre -> npp
## 14 3600 negative 0.12023142 pre -> npp
## 15 4900 negative 0.23867121 pre -> npp
## 16 6400 negative 0.30815296 pre -> npp
## 17 100 dark 0.06048052 pre -> npp
## 18 400 dark 0.11029673 pre -> npp
## 19 900 dark 0.16639601 pre -> npp
## 20 1600 dark 0.22696194 pre -> npp
## 21 2500 dark 0.27885166 pre -> npp
## 22 3600 dark 0.31852011 pre -> npp
## 23 4900 dark 0.35338378 pre -> npp
## 24 6400 dark 0.37870193 pre -> npp
## 25 100 positive 0.13945164 npp -> pre
## 26 400 positive 0.19473737 npp -> pre
## 27 900 positive 0.25254635 npp -> pre
## 28 1600 positive 0.31327780 npp -> pre
## 29 2500 positive 0.36524631 npp -> pre
## 30 3600 positive 0.41042253 npp -> pre
## 31 4900 positive 0.43152740 npp -> pre
## 32 6400 positive 0.45137261 npp -> pre
## 33 100 negative 0.16639620 npp -> pre
## 34 400 negative 0.24974175 npp -> pre
## 35 900 negative 0.49955110 npp -> pre
## 36 1600 negative 0.66653262 npp -> pre
## 37 2500 negative 0.83295414 npp -> pre
## 38 3600 negative 0.99940859 npp -> pre
## 39 4900 negative 0.78807485 npp -> pre
## 40 6400 negative 0.66641031 npp -> pre
## 41 100 dark 0.08290872 npp -> pre
## 42 400 dark 0.14873362 npp -> pre
## 43 900 dark 0.22827942 npp -> pre
## 44 1600 dark 0.28941655 npp -> pre
## 45 2500 dark 0.32906970 npp -> pre
## 46 3600 dark 0.36571275 npp -> pre
## 47 4900 dark 0.39113087 npp -> pre
## 48 6400 dark 0.41925841 npp -> pre
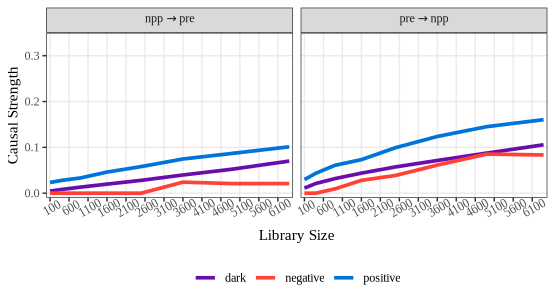
plot(npp_convergence, ylimits = c(-0.01,1.05),
xlimits = c(0,6500), xbreaks = seq(100, 6400, 500))